PanDDA is extensively explored by FragMAXapp. The webapp provides some options as interactive input. All options not available in the PanDDA tab is still available is parsed into the Custom parameters field.
Please check PanDDA documentation for all options available.
Data selection
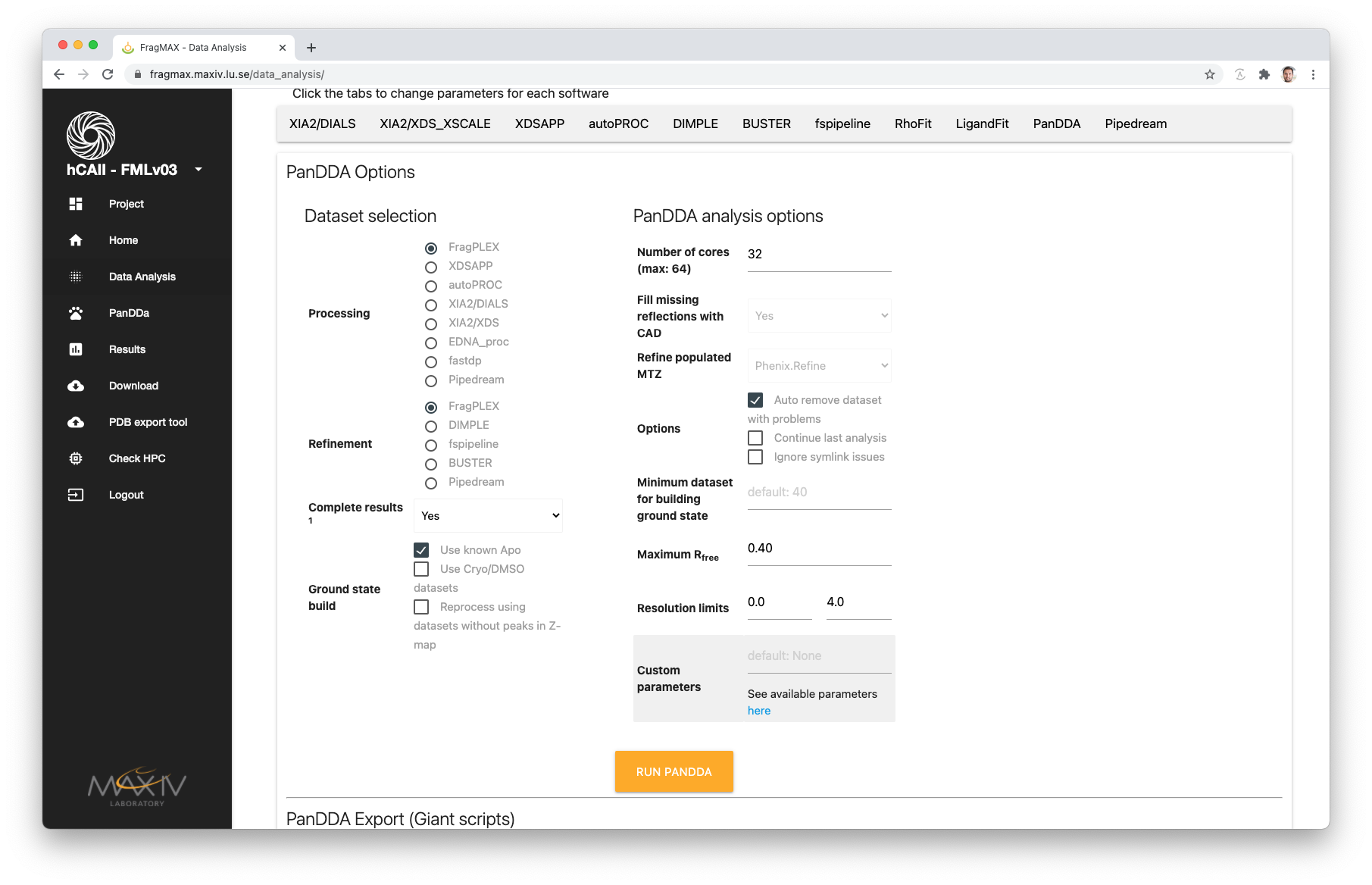
The selection will be used to find results in the project and analyse with PanDDA.
Selecting methods will primarily use the results from those pipelines. Example:
Processing = autoPROC
Refinement = DIMPLE
Output:
All results from the combination autoPROC processed and DIMPLE refined will be used.
If the a given dataset failied to be analysed by this combination, two options are possible:
1. If complete results = YES: FragPLEX method will be triggered,
looking for results availables from other method combinations.
2. If complete results = NO: Missing datasets will be ignored.
Options for generating the ground state model
Four options are available:
GSM = Ground State Model
1. Use know Apo structures
Datasets with _Apo_ in the name will be used to create the GSM
2. Use Cryo/DMSO datasets
Datasets marked as _opt_ or _DMSO_ will be used to create the GSM
3. Reprocess using dataset without peaks in Z-map
If a previous run is available, FragMAXapp will read the inspection log and use
all datasets without any events to build the GSM
4. Opening the dataset selection in the top of the page (the same for cheery pick
datasets for processing) and marking datasets will send the selection to PanDDA. This
option will replace options 1, 2 or 3. To ignore this behaviour, please make sure the
box _Process all datasets_ is marked.
NOTE: For all these options, if there is not enough datasets to create the GSM, the selection
will be ingored and PanDDA will define the datasets by itself.
Users can overwrite this behaviour by changing the minimum number of datasets to build GSM.
This option is availble in the middle colum (PanDDA analysis options)
PanDDA analyse

A slightly modified PanDDA HTML output is available, with options for checking the Run log and open the event maps
- Previous analysis are available in the tab, organised by date.
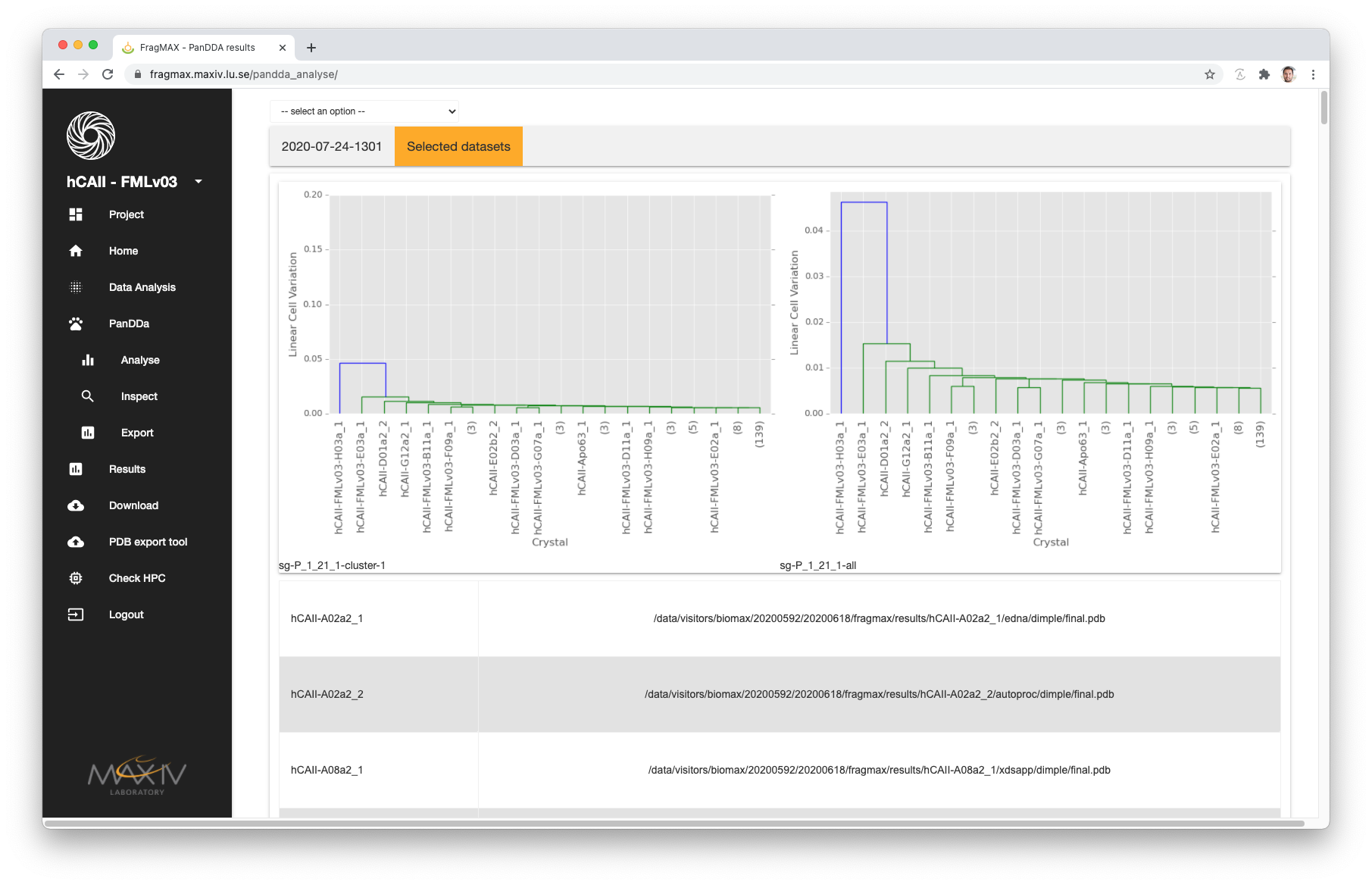
- To find the information from which method FragPLEX selected each dataset, the option Selected datasets is availble.
- In this tab, users will have a table with dataset name and path to the refined structure.
- A plot with the cluster analyse generated using Giant scripts
- A dropdown menu on the top left-hand side displays all different methods analysed by PanDDA
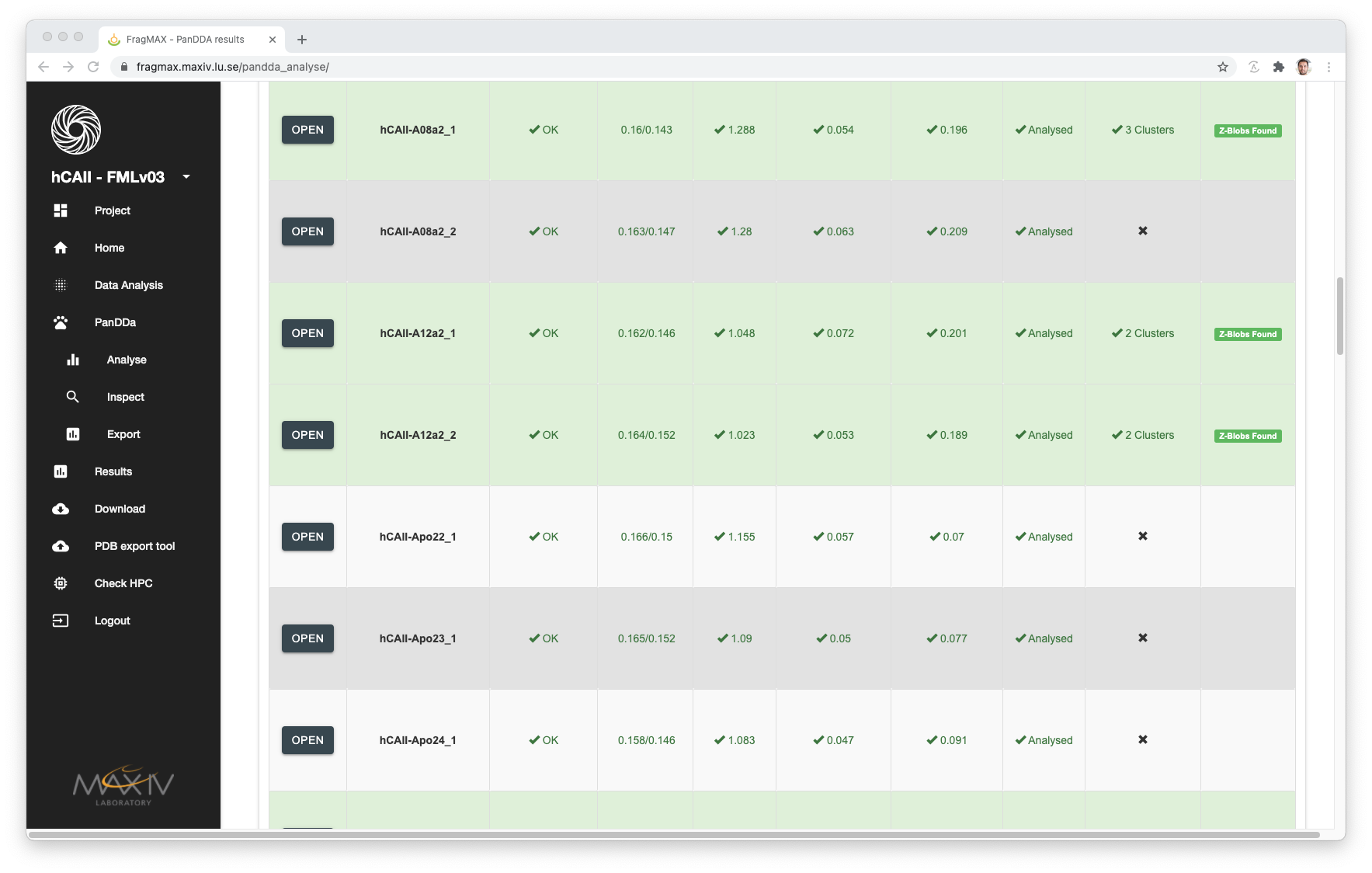
PanDDA inspections
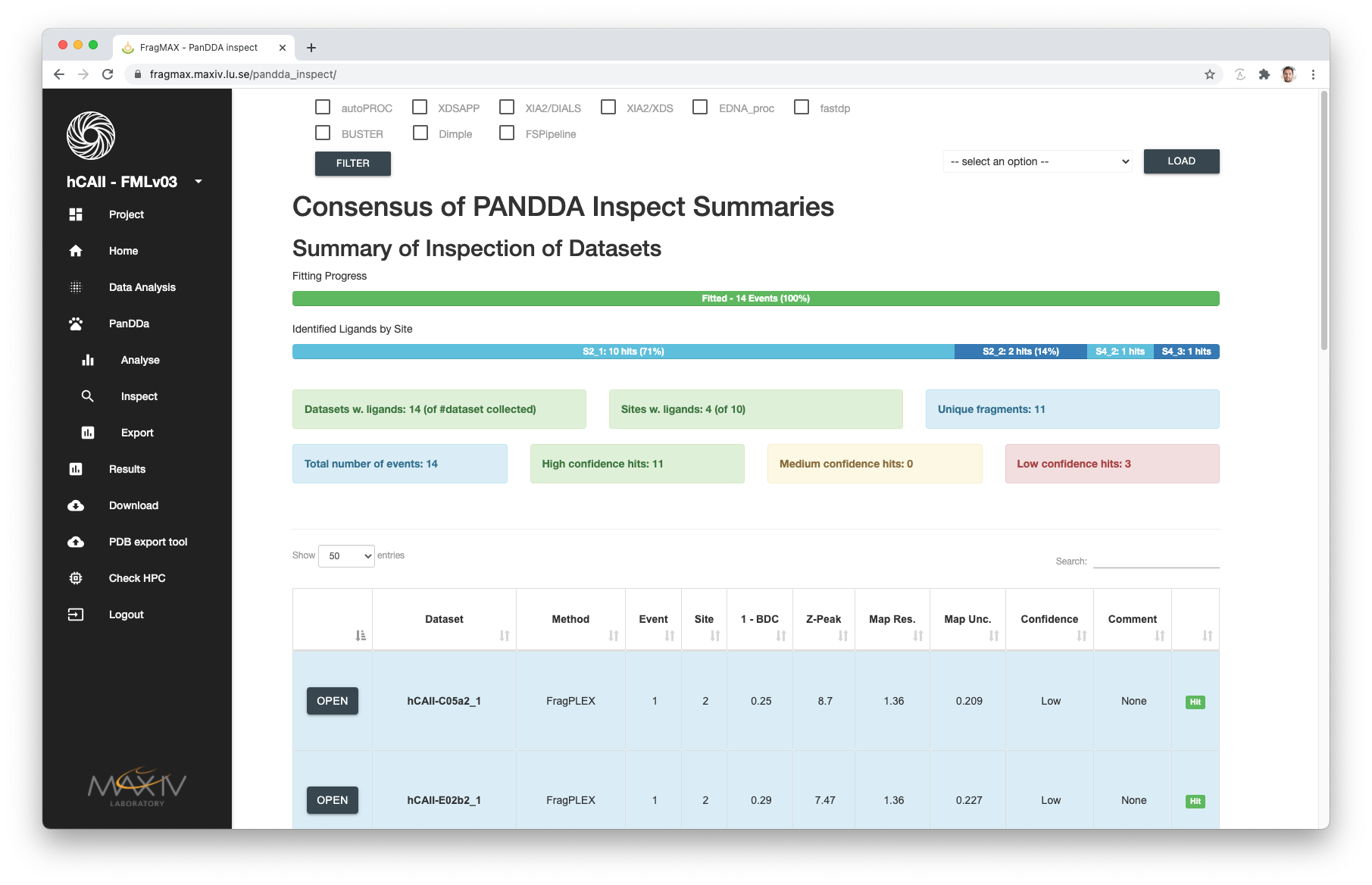
For analyses that was already inspected by the user and saved, the inspection HTML will be available.
If more than one inspection was made (with different Processing + Refinement selections), FragMAXapp will summarise the results in the consensus page.
- Loading individual inspections is possible using the dropdown menu in the right-hand side.
- Selecting the checkboxes on the top of the page and clicking filter will recreate the consensus summary using only those pipelines.
PanDDA event maps visualisation
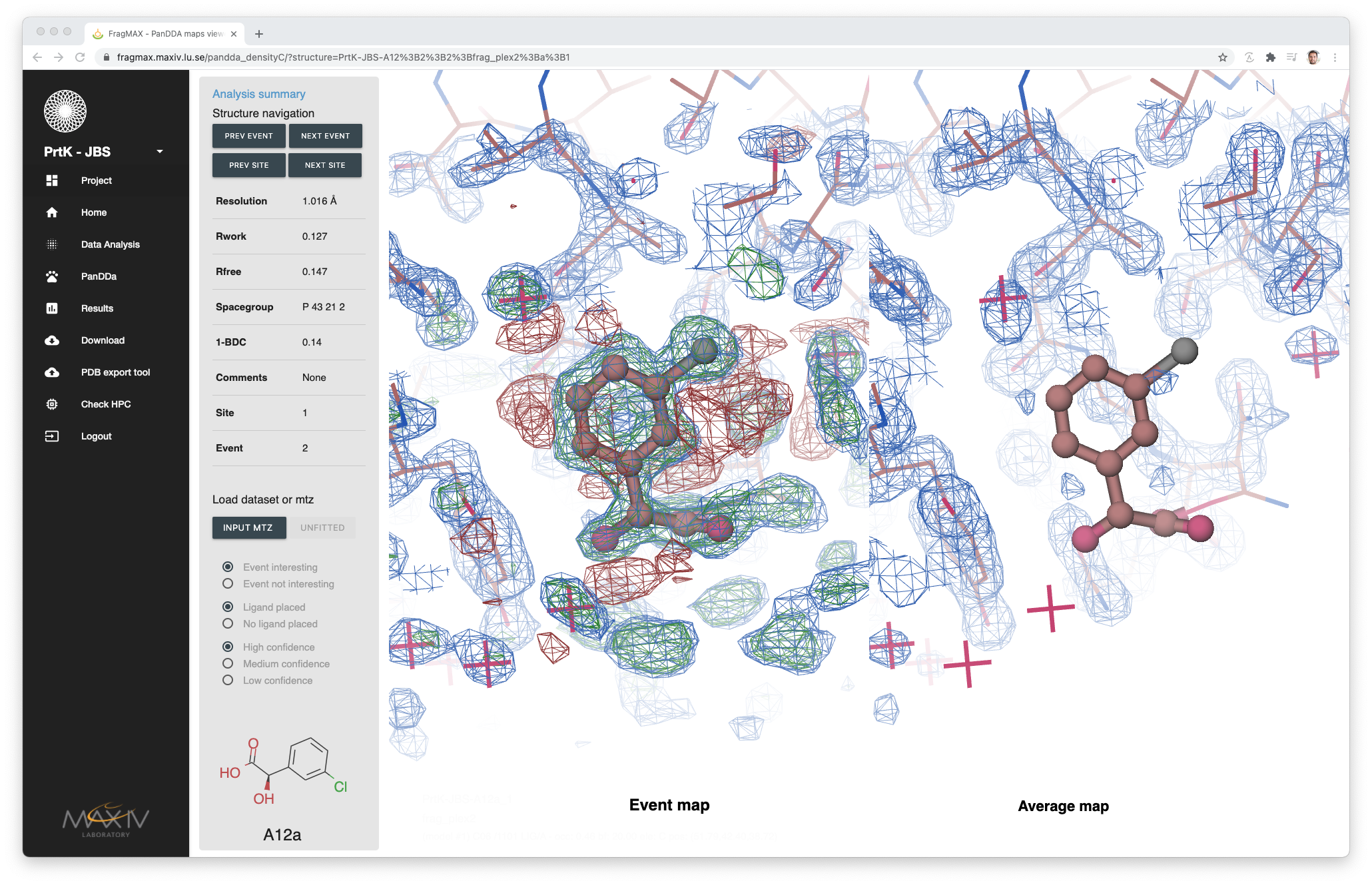
Opening the results from PanDDA will show a different interactive control bar than from regular results.
In this page, two mas will be loaded side-by-side: Event map and Average map. This is helpful to check if the density of the event is also present in the average model, reducing the amount of false positive results annotation.
- It is possible to load the input mtz (retrieved from the Structure Refinement method).
- PanDDA inspection annotation made during modelling is displayed and allows ammeneds.